 
Its wide variety of features are presented in an intuitive graphical user-interface, for which advanced computer skills are not required. However these products are no longer under development. QIAGEN CLC Main Workbench is used by tens of thousands of researchers all over the world for DNA, RNA and protein sequence data analysis. Hardware Įarly on, the company initially presented own-developed high-performance computing solutions, focusing on accelerating open source algorithms such as HMMER, Smith-Waterman and ClustalW, using FPGA technology. In 2020, CLC bio released a free plug-in that enables workflow execution on AWS directly from the CLC Genomics Workbench desktop software. The CLC Main Workbench was used for sequence analyses, including construction of phylogenetic trees using the MSAs and Neighbor Joining method. In 2019, this platform was adapted for and approved for use in AWS GovCloud. with packages ranging from basic (QIAGEN CLC Main Workbench). In 2017, CLC bio launched their CLC Genomics Cloud Engine as a command-line driven platform for cloud-based bioinformatics workflow execution on Amazon Web Services. On May 20, Cube Entertainment announced the official end of CLC, stating, Firstly. Features include read mapping and de novo assembly of high-throughput sequencing data, whole-genome detection of SNPs and structural variations, ChIP-seq, RNA-Seq, small RNA analysis, genome finishing, microbial genomics, structural biology, and functions to analyze, visualize, and compare genomic, transcriptomic, and epigenomic data. pathway analysis, genomics, and other omics). Īs additional capabilities were added to the software platform, it was eventually split into several themed Workbenches and plugins with collections of features relevant to different applications (e.g. In 2010, CLC bio was notable as the first commercial platform for bioinformatics analysis that utilized a graphical user interface for building, managing, and deploying analysis workflows as well as command-line tools, a SOAP and REST API, and later, the ability to run containerized tools. CLC bio developed some of their own open source algorithms, as well as their own SIMD-accelerated implementations of several existing popular applications. Software ĬLC bio's main activities were in software development for desktop ( Mac OS X, Windows, and Linux), enterprise, and cloud software for analysis of biological data. ĬLC bio was acquired by QIAGEN in 2013 and merged into its bioinformatics research and development division with several other purchased platforms in 2014. By 2012, it had additional offices in Cambridge, Massachusetts, Tokyo, Taipei and Delhi, with staff largely from research backgrounds (30% having a PhD) and had built a userbase of around 250,000 users in both academic institutions and biotechnology companies. Its product's development was also partly funded by collaborating with researchers on grant-funded projects. 
Error probabilities.CLC bio was a bioinformatics software company that developed a software suite subsequently purchased by QIAGEN.ĬLC bio started commercial activities on Januheadquartered in Aarhus, Denmark. CLC Main Workbench: Easy Access to DNA, RNA, and Protein Analyses Available for Windows, Mac OS X, and Linux platforms One hundred concurrent licenses. Learn more about QIAGEN CLC Genomics Workbench. Additionally, it includes all the sequence analysis tools of CLC Main Workbench. Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred. CLC Genomics Workbench is a comprehensive analysis package for the analysis and visualization of data and supports all typical NGS workflows.Ewing B, Green P (1998) Base-calling of automated sequencer traces using phred.Koch CM, Chiu SF, Akbarpour M, Bharat A, Ridge KM, Bartom ET, Winter DR (2018) A Beginner’s guide to analysis of RNA sequencing data.Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-seq.Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW (2016) Translating RNA sequencing into clinical diagnostics: opportunities and challenges.Ozsolak F, Milos PM (2011) RNA sequencing: advances, challenges and opportunities.Royce TE, Rozowsky JS, Gerstein MB (2007) Toward a universal microarray: prediction of gene expression through nearest-neighbor probe sequence identification. Genomes will be sequenced using the Illumina MiSeq system, de novo assemblies and gene annotations will be performed using the CLC Genomics Workbench.Okoniewski MJ, Miller CJ (2006) Hybridization interactions between probesets in short oligo microarrays lead to spurious correlations. 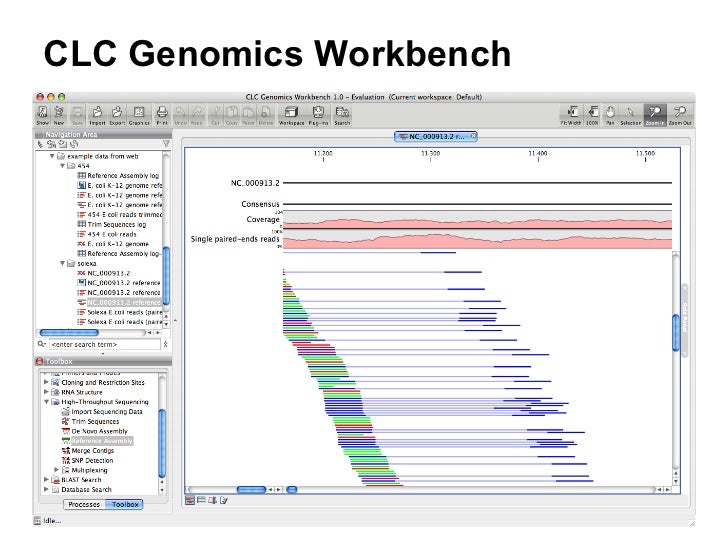
van Hal NL, Vorst O, van Houwelingen AM, Kok EJ, Peijnenburg A, Aharoni A, van Tunen AJ, Keijer J (2000) The application of DNA microarrays in gene expression analysis. 
Wang Z, Gerstein M, Snyder M (2009) RNA-seq: a revolutionary tool for transcriptomics. 
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
